Omicron has been derived from a well-known 'burst-type' search pipeline called Q-pipeline (a.k.a Omega). Q-pipeline is able to detect and characterize detector glitches with a very good efficiency and precision. So, the idea was to adapt it for detector characterization purposes, i.e. to run it over many auxiliary channels. To achieve this, the original matlab code has been replaced by a C++ architecture and many steps have been removed to run faster. The name Omicron was chosen to recall the Omega inheritance.
Index:
Data are loaded by chunk and chunks are analyzed by block (or segment). Segments and chunks overlaps with the previous iteration by an overlap defined by the user. The overlap must be large enough to avoid edge effects. The segment structure can be represented by this cartoon:
|________________chunk_________________|
| |_____________next chunk_____________|
|----seg----| |
|----seg----| |
|----seg----| |
|----seg----|
|--|
overlap
For each segment, triggers (t) are extracted as follows (overlap divided in 2):
|---------seg---------|
|ttttttttttttttt|
|
|---------seg---------|
|ttttttttttttttt|
The segment structure must be specified by the user in the option file. See Options file.
The power spectrum density (PSD) is computed following the median-mean method described in gr-qc/0509116. The PSD is estimated over one chunk of data. The PSD is estimated with a 0.5Hz resolution provided the search is conducted above 1Hz. For example if the duration of one chunk is 300s, the PSD is computed 300 times and the median-mean algorithm is applied to limit the bias due to loud glitches. The PSD is then interpolated to fit the channel sampling rate chosen by the user.
The PSD for each chunk of data can be saved for sanity checks. To do so, switch the TRIGGER/WRITEPSD option to 1.
To avoid dynamic range problems, the data is also high-pass filtered. This is done with zero phase distortion by first forward filtering and then reverse filtering the input data stream using a 12th order butterworth high pass filter. The cutoff frequency of the filter is specified in the option file by the lower edge of the frequency range.
This section summarizes the internal steps of Omicron. We used the example of a given event (glitch) measured by an accelerometer sensor. Here is the raw data time-series for this channel sampled at 8192 Hz. Only one second around the glitch of interest is plotted.
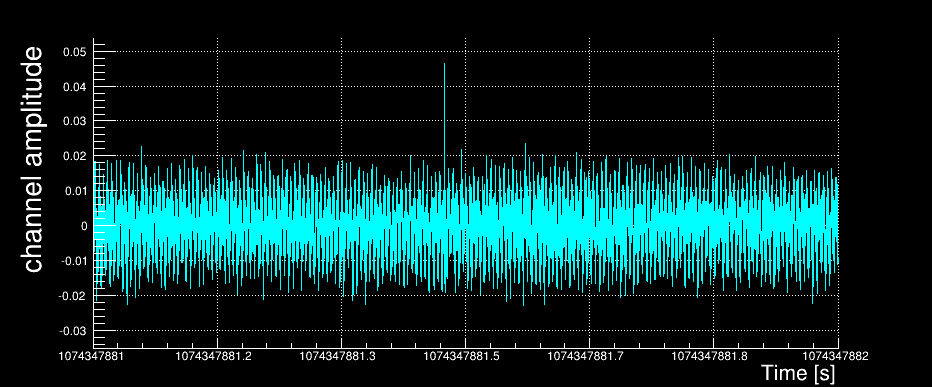
After being loaded, the data vector is down-sampled to a user-defined working frequency. In our example, we kept the channel native frequency of 8192 Hz.
The data vector is then high-passed at the lower range frequency value of the search (specified by the user). This is done with zero phase distortion by first forward filtering and then reverse filtering the input data stream using a 12th order butterworth high pass filter. In our example we used the search frequency range: 32Hz → 4096 Hz.
The power spectral density (PSD) is then computed using the median-mean method over the full time chunk. Here is the result for our example:
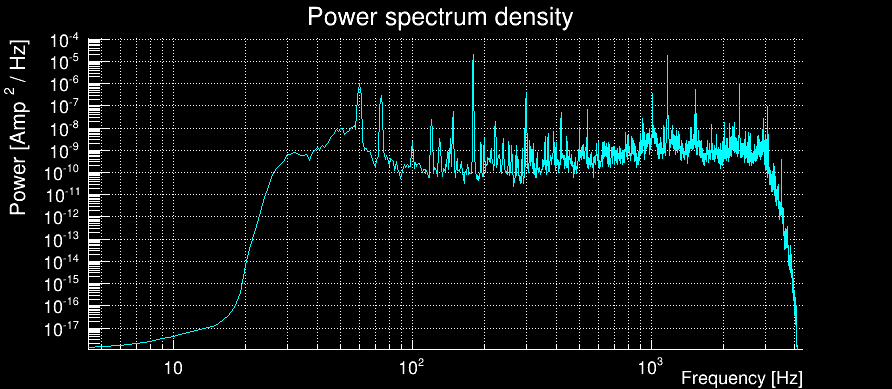
The 32Hz frequency cutoff is well visible.
The data vector is Fourier-transformed and normalized using the PSD. This data is projected over the Omicron tiles and the tile energy is computed. An amplitude SNR value is finally obtained for each tile. Here is what it looks like for our glitch (6 Q-planes were used and the vertical scale is the SNR):
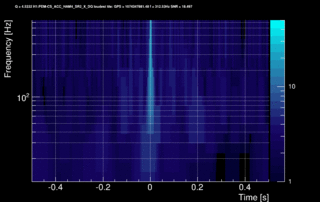
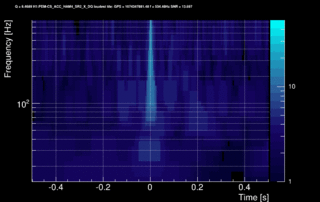
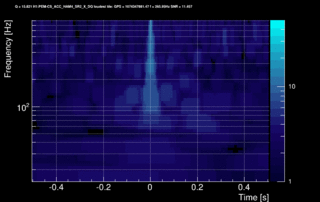
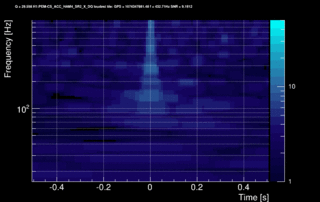
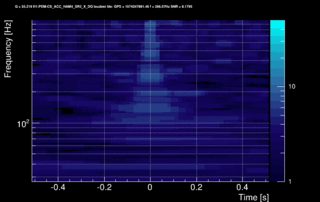
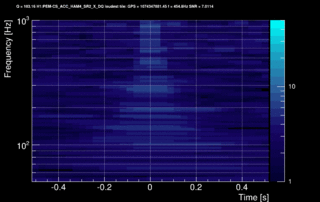
These 6 plots can be combined into one by overlapping the tiles and plotting the highest SNR tile on the top:
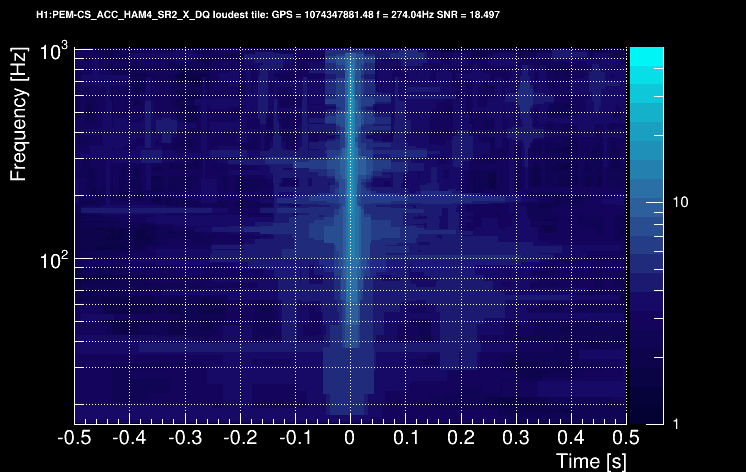
Only tiles with a SNR above a user-specified threshold are retained and saved on disk. The time and frequency information is given by the tile central point. An amplitude parameter is computed for each tile. It is given by SNR x sqrt(power) where power is a weighted average of the PSD over the tile.
Tiles are usually time-clustered to describe an event in totality. In this case, the cluster takes the time/frequency/SNR/amplitude value of the highest-SNR tile. In our example, the glitch is characterized by:
- time = 1074347881.482 s
- frequency = 274.04 Hz
- SNR = 18.497
- Amplitude = 1.55e-04 Amp. / sqrt(Hz)
The parameter space is tiled in 3 dimensions: Q, frequency and time.
First, a set of Q-planes is defined according to the maximum mismatch value specified by the user. Each Q-plane is then divided in frequency rows logarithmically distributed. Finally, each frequency row is divided in tiles linearly distributed over time.
The tiling structure only depends on the Q range , the frequency range, the segment duration and the working sampling frequency defined by the user. It it therefore built once for all when Omicron is launched.
A Omicron trigger is a tile with a signal-to-noise ratio above a given threshold.
Triggers produced by Omicron are stored in a TTree structure (named 'triggers') saved in a ROOT file, a XML file or dumped in a TXT file. The output file is created for each channel and the file name follows the convention:
[channel name]_[GPS first]_[duration].root (or .txt) [channel name] is the name of the processed channel [GPS first] is the GPS time of the first trigger [duration] is the duration in seconds between the first and last triggers
The triggers produced by Omicron follow the GWOLLUM convention. For mor details, see the GWOLLUM convention.
To get some help to directly read a ROOT trigger output file see How to read the Omicron output
It is also possible to change the output format in TXT or XML. The user must be aware that this choice is a dead-end for GWOLLUM. Indeed every GWOLLUM-based applications expect triggers to have a ROOT format with the GWOLLUM convention.
First you need to make sure your environment is well-defined by sourcing the Omicron setup script
Then, to run Omicron, you need to provide a timing and a parameter file:
omicron 993574004 993574123 ./parameters.txt
This command line runs Omicron between GPS=993574004 and GPS=993574123, using the parameters defined in ./parameters.txt. For the list of available options, please refer to Option file
Omicron can also be run over a segment file instead of a single time segment:
omicron ./segments.txt ./parameters.txt
where ./segments.txt is a text file with 2 columns [GPS start] [GPS end] following the GWOLLUM convention for segments.
The option file is composed of a list of keywords to define a given parameter. In the following we list all the available keywords. None of them are mandatory. If an option is not provided, a default value is assigned or is guessed from the available information
Click here to get an example of option file
→ DATA FFL [path to ffl file]
Path to the 'frame file list' file to access the desired channels.
Two formats are supported:
- the 'ffl' format (Virgo convention, 5 columns):
[frame file path] [GPS start] [file duration] [0] [0]
- the 'lalcache' format (LIGO convention, 5 columns):
[observatory] [frame type] [GPS start] [file duration] [frame file path]
The input format is automatically detected by Omicron
This option is not mandatory but it should be used for a standard Omicron processing.
→ DATA CHANNELS [list of channels]
List of main channels to process. Channel names must be separated by spaces or this option
can be used on several lines when listing channels
Default = 'V1:Pr_B1_ACp'
→ INJECTION CHANNELS [list of injection channels]
Injection channels can be added to the main channels.
There must be exactly as many injection channels as main channels.
→ INJECTION FACTORS [list of factors for injection channels]
Injection channels are added with factors provided with this option
There must be exactly as many factors as injection channels.
→ DATA SAMPLEFREQUENCY [sampling frequency]
Sampling frequency to which channels will be downsampled.
Upsampling is impossible.
Only one value is possible, common to all channels.
This parameter is very important since it will strongly influence the processing time.
Default = twice the upper end value of the frequency range
→ PARAMETER CHUNKDURATION [chunk duration]
→ PARAMETER BLOCKDURATION [block duration]
→ PARAMETER OVERLAPDURATION [overlap duration]
Chunk, block and overlap durations in seconds.
The three values of duration must verify the relation described in the segment section.
If not, a suggestion will be given when starting Omicron.
Default = 512
= 64
= 8
→ PARAMETER FREQUENCYRANGE [minimum frequency] [maximum frequency]
Frequency range for the search.
If [maximum frequency] is too large, the Nyquist frequency will be used
Default = 2 - sampling/2 Hz
→ PARAMETER QRANGE [minimum Q] [maximum Q]
Q range for the search. Q must be greater than sqrt(11).
Default = sqrt(11) - 100
→ PARAMETER FFTPLAN [FFT plan]
[FFT plan] = ESTIMATE, MEASURE, PATIENT is used to set the FFT plan of FFTW
Default = ESTIMATE
→ PARAMETER WINDOWS [list of time windows]
This option is only used when running omiscans. It lists the time windows (in s)
to be used for the time-frequency maps
Default = 2 8 32
→ PARAMETER MISMATCHMAX [maximum mismatch]
Maximum energy mismatch between tiles.
It must be a number between 0 and 0.5.
Default = 0.2
→ TRIGGER SNRTHRESHOLD [SNR threshold]
SNR threshold applied to each tile.
Default = 8
→ TRIGGER NMAX [maximum number triggers]
Maximum number of triggers per output root file.
Default = 1,000,000
→ TRIGGER RATEMAX [maximum trigger rate]
Alternatively, the maximum number of triggers per chunk can be set as a maximum rate:
NMAX = RATEMAX*CHUNKDURATION
→ TRIGGER CLUSTERING [clustering method]
Cluster triggers before saving them on disk.
Several clustering methods are available "TIME" and "Q"
If this option is not given, triggers are not clustered.
Default = no clustering
→ TRIGGER CLUSTERDT [δt parameter]
When the "TIME" clustering algorithm is selected,
it is possible to set the δt parameter with this option.
Default = 0.1s
→ OUTPUT DIRECTORY [path output directory]
Path to output directory.
The directory must be created before running Omicron.
All the output files of Omicron will be written in this directory.
Default = current directory
→ OUTPUT FORMAT [output file formats]
Output files can be saved with a root format (root), a XML format (xml)
and a text format (txt).
Additionally, image formats can be used (gif, png, jpg...) to save other outputs:
Omiscan maps, PSD...
The requested formats must be separated by white spaces
Default = root
→ OUTPUT PLOTSTYLE [plotting style]
Plotting styles supported by GwollumPlot: GWOLLUM, STANDARD and WHITEGWOLLUM
Default = GWOLLUM
→ OUTPUT VERBOSITY [verbosity level]
verbosity level: 0, 1, 2 or 3
0: no printing
1: basic printing
2: parameters printing
3: full printing (debugging)
Default = 1
→ OUTPUT WRITEPSD [0/1]
if 1, then write/plot the PSD of each chunk
Default = 0
→ OUTPUT WRITETIMESERIES [0/1]
if 1, then write/plot the data time series (after resampling) of each chunk
Default = 0
Omicron triggers can be printed with this tool. The command is very simple if triggers were centrally produced:
GetOmicronTriggers -c V1:h_4096Hz 934228815 934232415
If you produced your own triggers, you need to specify the trigger files location:
GetOmicronTriggers -c V1:h_4096Hz -t "/path/to/my/triggers/*.root" 934228815 934232415
This command prints every triggers between GPS times 934228815 and 934232415 for the channel 'V1:h_4096Hz'.
This tool is highly tunable. You can control the output, de-activate the clustering, select the triggers... For a complete list of available options, type:
GetOmicronTriggers -h
To get the list of available channels, type:
GetOmicronTriggers -l
IMPORTANT note: by default, triggers are time clustered. This takes quite some time. You can de-activate this option with the '-C NO' option.
This tool runs an OmiScan. The command is very simple if triggers were centrally produced:
GetOmicronScan -m V1:h_4096Hz -g 934232415
If you produced your own triggers, you need to specify the trigger directory:
GetOmicronScan -m V1:h_4096Hz -r /path/to/my/triggers/ -g 934232415
This command produces an OmiScan report of GPS=934232415 in the current directory. The option '-m' defines the name of the main channel (the one which is always plotted and is plotted first).
This tool is tunable. For a complete list of available options, type:
GetOmicronScan -h
Omicron trigger files (ROOT) can be printed with this tool. The command is very simple:
GetTriggerFileList -c V1:h_4096Hz 934228815 934232415
This command prints the trigger path which contain triggers between GPS times 934228815 and 934232415 for the channel 'V1:h_4096Hz'.
To monitor the triggers produced by Omicron, use this tool to print a set of plots (for centrally-produced triggers):
GetOmicronPlots -c V1:h_4096Hz -s 934228815 -e 934232415
If you produced your own triggers, you need to specify the trigger files location:
GetOmicronPlots -c V1:h_4096Hz -t "/path/to/my/triggers/*.root" -s 934228815 -e 934232415
This command plots the trigger properties of channel 'V1:h_4096Hz' between GPS times 934228815 and 934232415.
This tool is tunable. For a complete list of available options, type:
GetOmicronPlots -h
With this tool, the Omicron trigger plots built with GetOmicronPlots can be nicely presented on a web page. For example:
GetOmicronWeb -c V1:h_4096Hz 934228815 934232415
This tool is tunable. For a complete list of available options, type:
GetOmicronWeb -h
ROOT provides many functions to access and plot data. Here are a few basic examples obtained when working on a Omicron output file called 'V1:h_16384Hz_993571205_3351.root'
How to easily browse the data?
You can open a TBrowser and use the interface to navigate in the TTree called 'triggers':
florent-laptop:~> root V1:h_16384Hz_993571205_3351.root root [0] Attaching file V1:h_16384Hz_993571205_3351.root as _file0... root [1] TBrowser aAfter double-clicking on the triggers TTree, you will see all the variables defining your triggers. You can plot any of them by double-clicking on the 'leaves'. Remember the variable names, it will be helpful for the next examples.
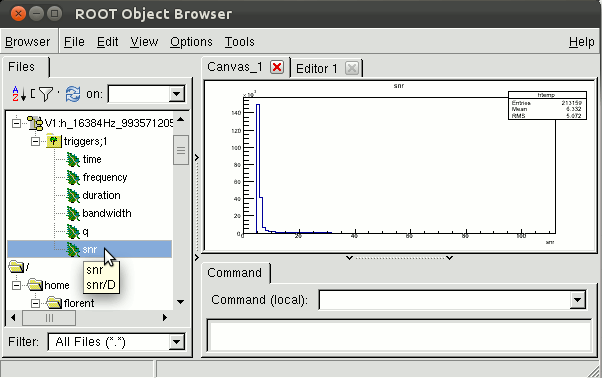
How to plot anything you want?
Any plot can be obtained in a single command line. For example:
florent-laptop:~> root V1:h_16384Hz_993571205_3351.root
root [0]
Attaching file V1:h_16384Hz_993571205_3351.root as _file0...
root [1] triggers->Draw("frequency:time")
will give you the time frequency map of your triggers. Right-click on the different parts of the canvas (frame, graph, axis,...) allows you to change the style of your plot (colors, scales, titles,...).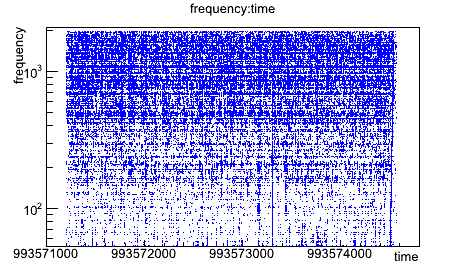
root [2] triggers->Draw("snr:time","frequency>100&&frequency<200")
will give you the SNR as a function of time for triggers with frequency between 100 and 200Hz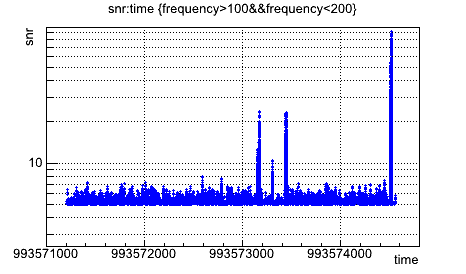
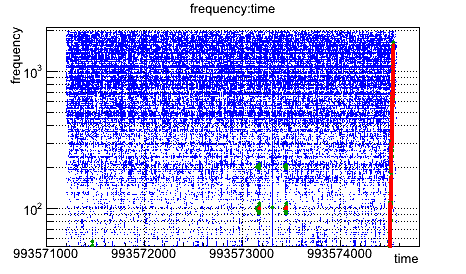
root [3] triggers->Draw("frequency:time")
root [4] triggers->Draw("frequency:time","snr>10","same")
root [5] triggers->Draw("frequency:time","snr>20","same")
In the same spirit, a glitchgram can be obtained by sumperimposing 3 graphs and fixing the colors.
root [6] triggers->Draw("log10(snr):(time-993571205)")
is a dummy example to show how to perform more advanced operations in the plotting. Here we shifted the time origin to 0 and we plotted the log10 of the SNRs.
How to print the data?
Open the ROOT file as before and use the 'Scan' function instead of 'Draw':
root [7] triggers->Scan("time:frequency:snr:log10(snr)","snr>20")
will print 4 columns listing the triggers with SNR>20.
All this is ofcourse just a glimpse of what is possible with ROOT, for more information go to the ROOT web page
|
|
Florent Robinet |